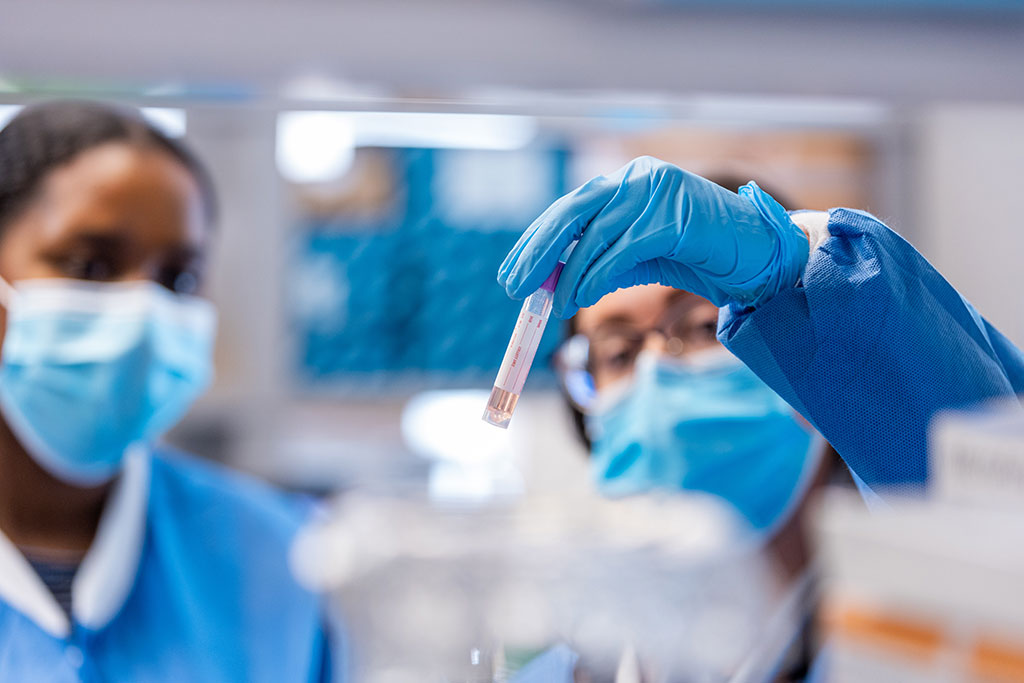
“You can’t fix what you don’t measure” is a maxim in the business world. And it holds true in the world of public health as well.
Early in the pandemic, the United States struggled to meet the demand to test people for SARS-CoV-2. That failure meant officials didn’t know the true number of people who had COVID-19. They were left to respond to the pandemic without knowing how quickly it was spreading and what interventions minimized risks.
Now the U.S. faces a similar issue with a different type of test: genetic sequencing. Unlike a COVID-19 test that diagnoses infection, genetic sequencing decodes the genome of SARS-CoV-2 virus in samples from patients. Knowing the genome sequence helps researchers understand two important things – how the virus is mutating into variants and how it’s traveling from person to person.
Before the COVID-19 pandemic, this kind of genomic surveillance was reserved mainly for conducting small studies of antibiotic-resistant bacteria, investigating outbreaks and monitoring influenza strains. As genomic epidemiologists and infectious disease experts, we perform these kinds of tests every day in our labs, working to puzzle out how the coronavirus is evolving and moving through the population.
Particularly now, as new coronavirus variants of concern continue to emerge, genomic surveillance has an important role to play in helping bring the pandemic under control.

Tracking Virus’s Travels and Changes
Genome sequencing involves deciphering the order of the nucleotide molecules that spell out a particular virus’s genetic code. For the coronavirus, that genome contains a string of around 30,000 nucleotides. Each time the virus replicates, errors are made. These mistakes in the genetic code are called mutations.
Most mutations do not significantly change the function of the virus. Others may be important, particularly when they encode vital elements, such as the coronavirus spike protein that acts as a key to enter human cells and cause infection. Spike mutations may influence how infectious the virus is, how severe the infection may become, and how well current vaccines protect against it.
Researchers are particularly on the lookout for any mutations that distinguish virus specimens from others or match known variants.
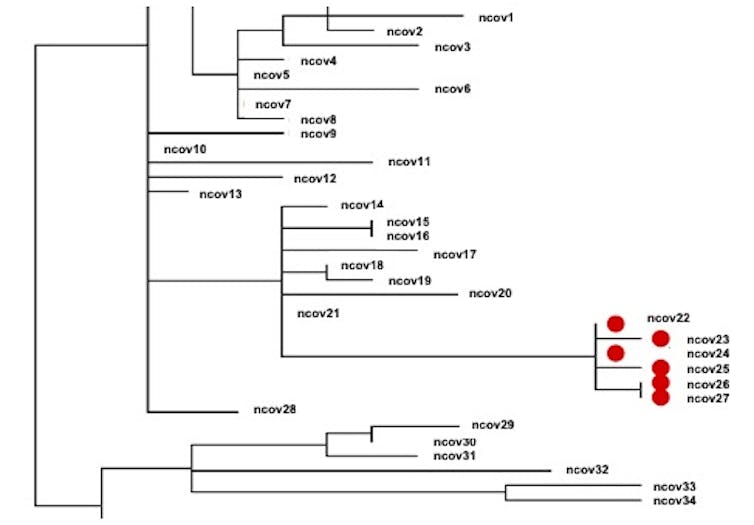
Scientists can use the genetic sequences to track how the virus is being transmitted in the community and in health care facilities. For example, if two people have viral sequences with zero or very few differences between them, it suggests the virus was transmitted from one to the other, or from a common source. On the other hand, if there are a lot of differences between the sequences, these two individuals did not catch the virus from each other.
This kind of information lets public health officials tailor interventions and recommendations for the public. Genomic surveillance can also be important in health care settings. Our hospital, for example, uses genomic surveillance to detect outbreaks that otherwise are missed by traditional methods.
Surveillance Can Provide a Warning
But how do researchers know if variants are emerging and if people should be concerned?
Take the B.1.1.7 variant, first detected in the United Kingdom, which has strong genomic surveillance in place. Public health investigators discovered that a certain sequence with multiple changes, including the spike protein, was on the rise in the U.K. Even amid a national shutdown, this version of the virus was spreading rapidly, more so than its predecessors.
Scientists looked further into this variant’s genome to determine how it was overcoming the distancing recommendations and other public health interventions. They found particular mutations in the spike protein – with names like ∆69-70 and N501Y – that made it easier for the virus to infect human cells. Preliminary research suggests these mutations translated into a higher rate of transmission, meaning that they spread much more easily from person to person than prior strains.
Vaccine developers and other scientists then used this genetic information to test whether the new variants change how well the vaccines work. Fortunately, preliminary research that has not yet been peer-reviewed found that the B.1.1.7 variant remains susceptible to current vaccines. More worrisome are other variants such as P.1. and B.1.351, first discovered in Brazil and South Africa, respectively, that can evade some antibodies produced by the vaccines.
Setting Up a Genomic Surveillance System
Detecting variants of concern and developing a public health response to them requires a robust genomic surveillance program. That translates to scientists sequencing virus samples from about 5% of the total number of COVID-19 patients, selected to be representative of the populations most at risk from the disease. Without this genomic information, new variants may spread rampantly and undetected through the country and globally.
So how is the U.S. performing in the area of genomic surveillance? Not very well, and well behind other developed countries, coming in 34th in the number of SARS-CoV-2 genomes sequenced per number of cases. Even within the U.S., there is large variation among states for genomes sequenced per number of cases, ranging from Tennessee at 0.09% to Wyoming at 5.82%.
But this is about to change. The Centers for Disease Control and Prevention, in conjunction with other agencies of the federal government, is partnering with private labs, state and local public health labs, academia and others to increase genomic surveillance capacity in the U.S.

Reaching the new national goal of 5% set by the White House is not as simple as footing a hefty bill for a laboratory to perform the tests, though. Laboratories must collect the samples, often from different sources: public health labs, hospitals, clinics, private testing labs. Once the sequencing test is performed, bioinformaticians use advanced programs to identify important mutations. Next, public health professionals merge the genomic data with the epidemiological data to determine how the virus is spreading. All of this requires investment in training people to perform these tasks as a team.
Ultimately, to be useful, a successful genomic surveillance program must be fast and the data needs to be made publicly available immediately to inform real-time decision-making by public health officials and vaccine manufacturers. Such a program is one of the public health tools that will help bring the current pandemic under control and set up the U.S. to be able to respond to future pandemics.
ABOUT THE AUTHORS
Alexander Sundermann, MPH, CIC is a Clinical Research Coordinator and Doctor of Public Health (DrPH) student in Epidemiology at the University of Pittsburgh. Alex is part the Microbial Genomic Epidemiology Laboratory (MiGEL) that is creating an outbreak detection tool that will enable healthcare systems to detect, intervene, and stop outbreaks from occurring. The tool, Enhanced Detection System for Healthcare Association Transmission (EDS-HAT), combines machine learning with the electronic health record and has success in initial prospective findings. Alex has published manuscripts on novel healthcare-related outbreaks using MiGEL’s methods. Alex has prior experience as an Infection Preventionist at the University of Pittsburgh Medical Center where he also published on various quality improvement projects including mold contaminated linens.
Lee Harrison is Professor of Epidemiology, Medicine, and Infectious Diseases and Microbiology, University of Pittsburgh. Dr. Harrison is an infectious diseases physician whose research focuses on the epidemiology and genomic epidemiology of serious infectious pathogens, including S. pneumoniae, N. meningitidis, C. difficile, influenza and SAR-CoV-2. A current project involves novel genomic and computational methods for early detection of outbreaks of bacterial and viral pathogens in the hospital. This approach uses whole genome sequencing surveillance and data mining of the electronic health record using machine learning to identify serious outbreaks that are missed by traditional hospital epidemiologic methods.
Vaughn Cooper is the EvolvingSTEM Founder and Executive Director and a Professor of Microbiology and Molecular Genetics, University of Pittsburgh. He co-founded and is Director of the Center for Evolutionary Biology and Medicine (CEBaM), founded and directs the Microbial Genomics Sequencing center (MiGS) and is the Chair of the Council for Microbial Sciences (COMS) for the American Society of Microbiology. The Cooper laboratory studies how bacterial populations (e.g. Burkholderia, Pseudomonas, Acinetobacter, and multispecies communities) evolve to adapt to new hosts and environments, particularly in biofilms. Other major interests include the evolution of antimicrobial resistance and why genome regions mutate/evolve at different rates.
This article is courtesy of The Conversation.