Before the age of mass travel, infectious diseases went from local outbreak to international epidemic by ship. In the 21st century, global air travel, high speed trains, public transportation, and improved road systems, provide new means for diseases to spread internationally at unprecedented rates.
The world recently faced these public health disasters with the 2003 Severe Acute Respiratory Syndrome that killed over 800 people across 37 countries, the 2009 Influenza H1N1 which affected over 200 million individuals across the globe, and the 2014 Ebola that has killed over 11,000 people.
In all these cases, government officials scrambled to put response plans in place; however, by the time travel alerts and health recommendations were announced, the diseases had already reached epidemic proportions, costing thousands of lives and billions of dollars. It is incumbent on scientists to develop tools that will track and predict disease spread and identify interventions that can be executed in a timely and effective manner.
One such tool is the Spatiotemporal Epidemiological Modeler (STEM). Building on the open-source platform provided by the Eclipse Foundation, STEM is designed to analyze the global spread of infectious diseases and model the impact of public health interventions.
The unique nature of STEM is that it is a multi-disciplinary, collaborative modeling platform. The open-source characteristics of the system allow researchers and programmers to add, compare, refine, and validate different scenarios as well as add denominator data based on specialty. For example, an infectious disease specialist in dengue working in South America may have unique disease characteristics and population data that could be tailored into STEM.
Every component of STEM is an Eclipse “Plugin” which means that each part of STEM can be independently developed, exchanged, extended, added, or removed without having to re-write computer code for the entire program. Each customized section plugs right back into the system. The more professions that add data, models, and features, the stronger the tool becomes.
The disease models can be configured by known characteristics including infection rates, incubation period, and mortality rates. These variables can also be changed during the course of the scenario to depict possible mutations in the pathogen. At a more complex level, the vectors of disease, such as motion of individuals, ground travel, air travel, and social gatherings, that affect the transmission of an infectious disease can also be modeled.
STEM uses a compartment method of mathematical modeling to illustrate the spread of a disease using known disease parameters and environmental factors while allowing for the control of a host of other variables. A typical compartment model is the SEIR model where each person in a population, in relation to a particular disease, will be either in a state of: Susceptibility (S), Exposed (E), Infectious (I), or Recovered (R). This type of model assumes that members of the population will be in separate compartments; the same person will not be in different compartments at the same time. Transitions between compartments are driven by differential equations.
When new models are required, STEM provides a graphical model builder that allows users to define new compartment models (and the corresponding equations) without the need to write any code. The new plugins are automatically generated by the graphical design tool and instantly available. Compartments may be added or subtracted to match the normal characteristics of the disease. STEM provides drop-down menus so users can “plug and play” with different disease characteristics using the most appropriate compartment model.
Figure 1 show the screen of a typical SEIR model. Researchers can also edit the software to customize the models to fit any scenario. These new scenarios can then be shared with the rest of the community to continue the open-source collaboration.
STEM is pre-loaded with data on countries, national borders, transportation networks, air travel, and environmental conditions. Researchers can add their own data sources and disease characteristics to match their own particular situation. STEM can also be used to create customized graphs. This feature makes it possible, for example, to create a spatial map of animal pens on a farm and to import that graph into the model to study the spread of a veterinary disease.
STEM can also be used to model the effectiveness of public health interventions such as social distancing, vaccination, quarantine, and evacuation. STEM can provide researchers with data such as the incidence of the disease, the number infected, the number recovered, and fatality rates.
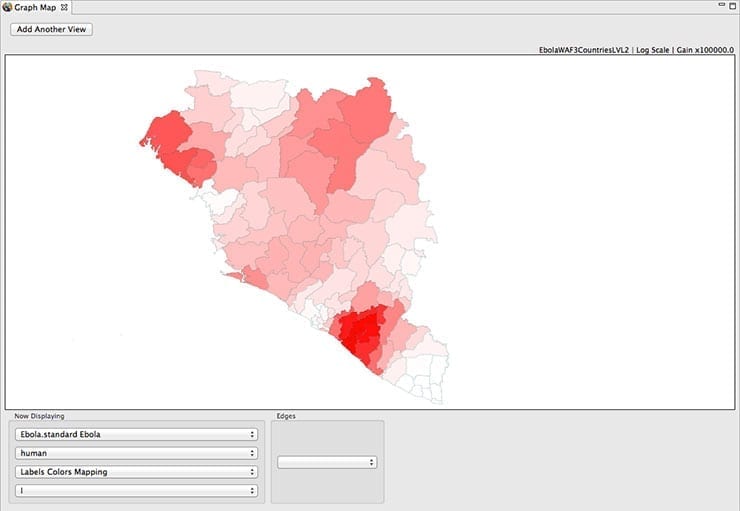
STEM contains several tools for analyzing this data including a statistical tool for comparing models and a data visualizer that can create maps of the outbreak. Figure 2 illustrates an example of an Ebola model showing a Guinea, Liberia, and Sierra Leone spread of disease based on the 2014 outbreak. The data can also be exported for analysis using other programs.
HOW TO USE STEM
STEM can be downloaded to any computer running Windows, Mac OS X, or other operating systems. The STEM website offers full download links, tutorials, installation guides, and YouTube video tutorials in English, Hebrew, Japanese, and Spanish.
STEM also has many sample projects that researchers from around the world have shared and made available for download. These scenarios include the recent 2014 Ebola epidemic, dengue fever, avian influenza, malaria, H1N1, and food-borne diseases. Other scenarios are part of tutorials that demonstrate basic functions of the tool. The STEM wiki page has a full list of STEM resources and publications that showcase disease models.
INTERESTED IN THE STEM COMMUNITY
Anyone can join the STEM community and participate in the monthly phone call. All are welcome to contribute to STEM, report bugs, or request new features or scenarios.
ACKNOWLEDGEMENTS
STEM is an Eclipse project originally created by IBM Research with contributors including the Federal Institute for Risk Assessment (BfR) in Germany, the Johns Hopkins Bloomberg School of Public Health, and many others. The STEM open-source community can be found at the project website: https://www.eclipse.org/stem/ and wiki: http://wiki.eclipse.org/index.php/STEM. Special thanks to Judith Douglas who contributed to the article as part of the STEM IBM project team.
DISCLAIMER
Development of STEM was supported, in part by the U.S. Air Force Surgeon General’s Office (USAF/SG) and administered by the Air Force District of Washington (AFDW) under Contract Number FA7014-07-C-0004. Neither Eclipse, the United States Airforce, IBM, nor any of their employees, nor any contributors to STEM, make any warranty, express or implied, or assumes any legal liability or responsibility for the accuracy, completeness, or usefulness of the STEM data sets. The Air Force has not accepted the products depicted and issuance of a contract does not constitute Federal endorsement of the IBM Almaden Research Center.
Contributing author Nereyda Sevilla is a Biodefense PhD student in in the School of Policy, Government, and International Affairs at George Mason University in Fairfax, VA (nsevilla@gmu.edu). Her dissertation uses STEM to examine the role of the global aviation system as an incubator and vector for infectious disease outbreaks. She is a proud member of the STEM community.